Archives
- 2026-04
- 2026-03
- 2026-02
- 2026-01
- 2025-12
- 2025-11
- 2025-10
- 2025-09
- 2025-03
- 2025-02
- 2025-01
- 2024-12
- 2024-11
- 2024-10
- 2024-09
- 2024-08
- 2024-07
- 2024-06
- 2024-05
- 2024-04
- 2024-03
- 2024-02
- 2024-01
- 2023-12
- 2023-11
- 2023-10
- 2023-09
- 2023-08
- 2023-07
- 2023-06
- 2023-05
- 2023-04
- 2023-03
- 2023-02
- 2023-01
- 2022-12
- 2022-11
- 2022-10
- 2022-09
- 2022-08
- 2022-07
- 2022-06
- 2022-05
- 2022-04
- 2022-03
- 2022-02
- 2022-01
- 2021-12
- 2021-11
- 2021-10
- 2021-09
- 2021-08
- 2021-07
- 2021-06
- 2021-05
- 2021-04
- 2021-03
- 2021-02
- 2021-01
- 2020-12
- 2020-11
- 2020-10
- 2020-09
- 2020-08
- 2020-07
- 2020-06
- 2020-05
- 2020-04
- 2020-03
- 2020-02
- 2020-01
- 2019-12
- 2019-11
- 2019-10
- 2019-09
- 2019-08
- 2019-07
- 2019-06
- 2019-05
- 2019-04
- 2018-07
-
It is of interest that
2021-04-19
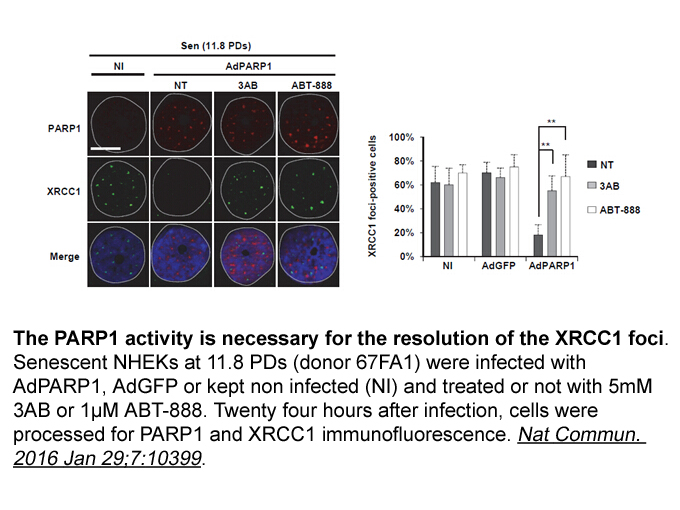
It is of interest that an initial maternal experience in virgin mice produces both changes in subsequent maternal behavior as well as possible changes in neural gene expression. Stolzenberg et al. (2012) proposed that experience-based changes in maternal responsiveness may be mediated by chromatin m
-
NF B is a family of protein
2021-04-19
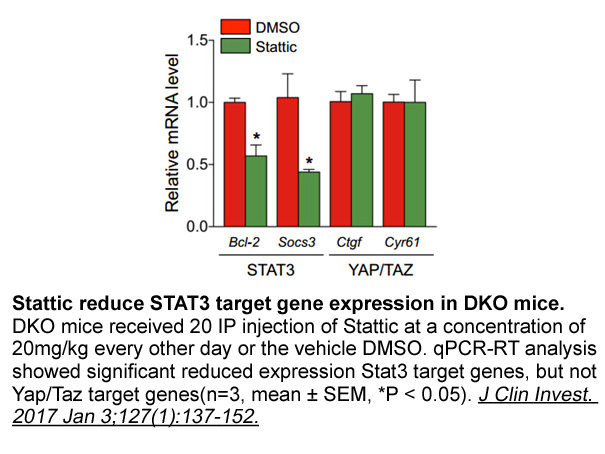
NF-κB is a family of protein mediators that regulate various innate and adaptive immune responses [29,30]. The NF-κB family consists of the following five proteins: c-Rel (Rel); p65 (RelA); RelB; p50(NF-κB1); and p52(NF-κB2). It has been confirmed that NF-κB is activated by TNF family cytokines, suc
-
In this work we fabricated a
2021-04-19

In this work, we fabricated a kind of multifunctional nanoparticles, based on HAuNS coated by chitosan-stearic vasopressin receptor antagonist copolymer (CSO-SA), which was employed frequently to form micelles for hydrophobic drug delivery in our previous work [21], [22]. The nanoparticles can enca
-
The interferences between thapsigargin and forskolin induced
2021-04-19
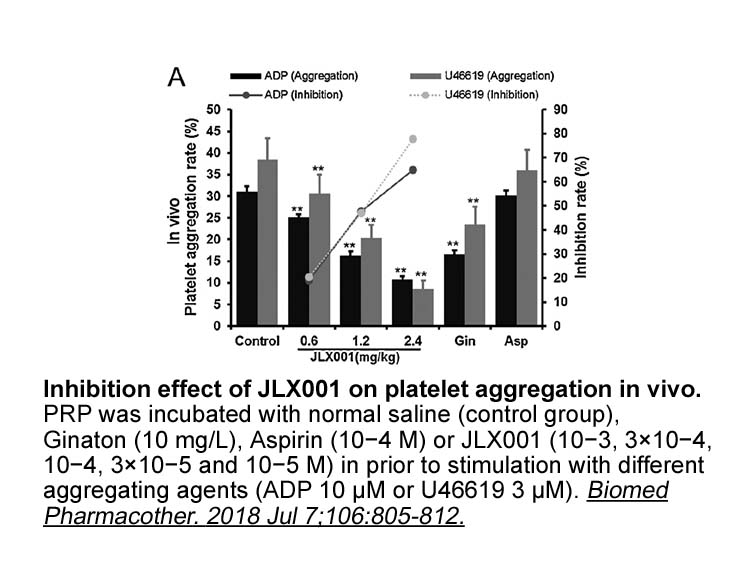
The interferences between thapsigargin- and forskolin-induced Ca release indicate that these drugs deplete the same intracellular stores in RASMC. In fact, after partial depletion of thapsigargin-sensitive stores, the forskolin-induced increases in [Ca]c were significantly reduced. Similarly, a prev
-
We next turned our attention
2021-04-19
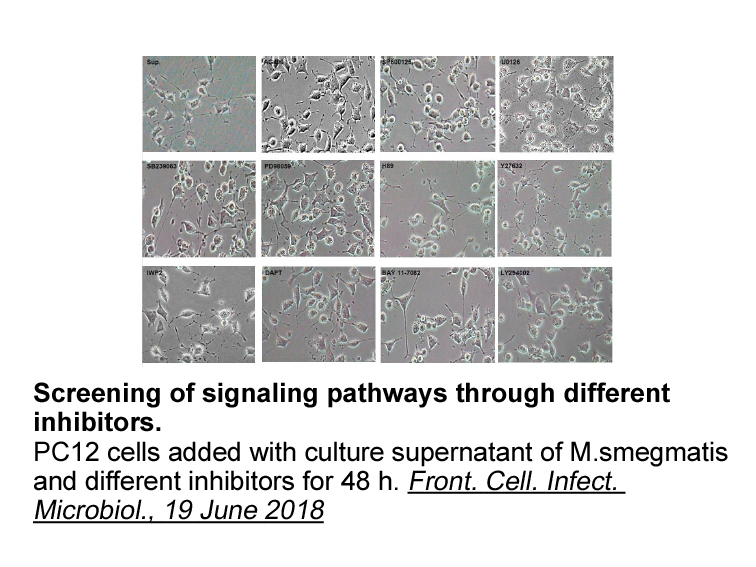
We next turned our attention to the 6-position of the indole (). It was found that the chlorine atom present in was extremely important. The des-chloro analogue showed nearly 30-fold less affinity for the EP receptor in the binding assay and was nearly 30-fold lower in activity in the functional a
-
Docking experiments showed that Me MeGlcA Xyl and MeGlc
2021-04-19

Docking experiments showed that Me-MeGlcA3Xyl4 and MeGlc3Xyl4 were bound to EcXyn30A and the R293A variant in the same way as MeGlcA3Xyl4, having MeGlcA or its modified forms accommodated in the -2b subsite. All ligands were coordinated by the same atomoxetine hcl and no new interactions were obser
-
Introduction Cholesterol plays a pivotal role as a constitue
2021-04-19
Introduction Cholesterol plays a pivotal role as a constituent of biological membranes and as a precursor for vitamins, hormones and bile acids. Accordingly, its production, distribution, and elimination must be tightly regulated at the cellular and organismal level. Accordingly, dysregulated chole
-
br Introduction Vitis vinifera L is one of the
2021-04-19
Introduction Vitis vinifera L. is one of the most worldwide-grown fruit crop and is mostly used in the wine industry (Bouquet, 2011). Domestication and history changed dramatically the biology of this species and today there are thousands of V. vinifera varieties, being many of them closely relat
-
Moreover maintaining the pyrimidine pool is not the only imp
2021-04-19
Moreover, maintaining the pyrimidine pool is not the only important role played by DHODHs. In Toxoplasma gondii, for instance, DHODH plays a second essential function [38] possibly coupled to the mitochondrial respiratory activity, where it replenishes ubiquinol levels and prevents reactive oxygen s
-
br Phylogenetic analysis KSTDs have
2021-04-19
Phylogenetic analysis — Δ1-KSTDs have rather diverse amino GSK J1 sequences. A sequence distance analysis using the program MEGA6 [75] of all currently biochemically characterized Δ1-KSTDs yielded a largest p-distance [76] of 0.67 (on a scale of 0-1) for the Δ1-KSTDs from the actinobacteria Nocar
-
Calphostin C mg br Methods br Results br Discussion
2021-04-17
Methods Results Discussion On the other hand, surgical patients have to some extent stress during pre- and/or post-operative period. It is well known that CRF/CRF1 receptor is involved in the activation of HPA during acute and chronic stress (Papadimitriou and Priftis, 2009, Zelena et al.,
-
It has been shown that CK phosphorylates and
2021-04-17
It has been shown that CK1δ phosphorylates α-, β- and γ-tubulin in vitro and that CK1δ specifically interacts with the trans Golgi network, COPI positive vesicles, and centrosomes in interphase cells [11], [12], [13], [14]. Moreover CK1δ is also associated with granular particles that are associated
-
Since clinical studies mostly involved cases that used
2021-04-17
Since clinical studies mostly involved cases that used MPA, data from studies that examined other types of progestogens are very limited. While the WHI study examined the combination of conjugated equine estrogens plus MPA [1], the Million Women Study [2] that also demonstrated an increased risk of
-
br Introduction Receptor tyrosine kinases
2021-04-17
Introduction Receptor tyrosine kinases (RTKs) are critically involved in the development and progression of human cancers and are therefore useful targets for anti-cancer therapies [1]. The Eph receptors represent the largest subfamily of receptor protein kinases and interact with ligands called
-
During the course of enzymatic digestion
2021-04-17
During the course of enzymatic digestion, protein is digested into large peptides and these large peptides are then further digested into small fragments. αs1-Casein f15–39, which contains 25 amino enfuvirtide mg residues was the longest iTRAQ-labelled peptide identified in the samples digested wit
16236 records 702/1083 page Previous Next First page 上5页 701702703704705 下5页 Last page